
CT temporal subtraction: techniques and clinical applications
Computed tomography (CT) has been the main noninvasive diagnostic technique for evaluating lung lesions. Although multidetector CT (MDCT) enables simultaneous increased z-axis coverage and thinner slice collimation, large numbers of axial images are generated by MDCT, which leads to reviewer fatigue during interpretation (1). Human perceptual errors, therefore, currently seem to be one of the most significant limiting factors in the detection of small lung lesions (2). Bone is one of the most common sites for metastasis in cancer along with lung. Although CT is a routine imaging modality to survey many types of cancer, bone metastases are often missed at CT because of their subtle findings. Other modalities, such as bone scintigraphy and positron emission tomography (PET), are useful to detect bone metastasis, but they still must be identified at CT anatomically.
In order to solve these problems, computer aided diagnosis (CAD) systems have attracted attention in recent years (3). Improvement of image interpretation speed and image analysis accuracy is expected by using the CAD system because it can reduce burden on interpreting physicians and reduce variation in diagnostic accuracy. Temporal subtraction (TS), which is one of the computer-aided detection (CADe) techniques, can remove most of the normal structures, such as blood vessels, ribs, muscles, by performing subtraction calculation processing between the current and the previous images of the same subject (4). CT-TS technique can enhance the subtle change between the CT images. Therefore, it has been developed for CADe of small or ground-glass lung nodules and small or faint bone metastases. In available CT-CAD system lesion candidates are usually indicated by symbol/mark on a computer-output image, however, in the TS system observers refer to the subtraction image without symbol/mark and then judge whether a “lesion” exist or not. Therefore, diagnostic performance is heavily affected by the image quality of the subtraction image. To obtain a high-quality subtraction image without subtraction artifacts, image registration is a key technology. Up to date, there are many rigid and non-rigid image registration techniques. If the image warping is incorrect, normal structures remain as the artifacts on the subtraction image, and as a result the image quality can be degraded. Many image warping techniques have been developed based on 2-dimensional (2D) images using a chest radiography (5-7), and TS on plain radiograph have already been commercially available. In general, CT images which is obtained different time series have temporal changes such as shape, size and location. A TS image on thoracic CT, which is obtained by subtraction of a previous image from a current one, has a 3-dimensional (3D) structure and the deformation of the subject in the axial direction needs to be considered 3D registration technique is necessary. Since early 2000, several attempts have been made to develop an image warping technique based on 3D images using thoracic MDCT images (8,9). Although the quality of the subtraction images based on 3D was relatively good in general, misregistration still appeared as artifacts on the subtraction images. It is, however, necessary to employ a more complex 3D registration, since the TS image obtained from two successive CT scans can have noticeable subtraction artifacts on very small lung structures.
Non-rigid image registration method for MDCT images
Image registration is an image processing technique to align two or more images of the same scene such as remote sensing (10-12). The techniques are widely used in medical image processing fields. Image registration method can be categorized into two types: (I) feature-based registration (13-15), and (II) image-based registration (16-23).
In feature-based methods, it is necessary to detect landmarks and features required for registration from images. The existence of these landmarks and features corresponding to the two images and the guarantee of their detection accuracy are the premise of the feature-based registration. The features for the registration include point cloud, edge and surface of the image. Ko et al. proposed a system for deification of nodules at the chest CT image (13). Besl et al. proposed a method for registration of 3D shape based on feature-points (14). Maintz et al. proposed a registration of CT and MR brain images based on correlation of image features, notably edge and ridge features (15). On the other hand, Rueckert et al. proposed a nonrigid registration image matching method based on free-form deformations and its application to MR imaging (20). Pariaswamy et al. converted locally affine but globally smooth images in their general purpose registration method (21,22). Walimbe et al. demonstrated registration of clinical CT and PET images using a interpolation method of 3D rotations and translations from discrete regid-body transformations (23).
In medical imaging, the techniques enable us to combine data from multiple modalities in a multiple time series. In recent years, new image registration frameworks based on convolutional neural networks (CNN) are proposed (24-26). Wu et al. (24) develop a learning-based image registration framework based on deep learning techniques. To identify intrinsic deep feature representations in image patches, they introduced the feature selection method using a convolutional stacked autoencoder. Hu et al. proposed a weakly-supervised CNN for multimodal image registration framework from a MR image (25). Eppenhof et al. proposed a fast registration method to estimate a transformation model based on 3D CNN (26).
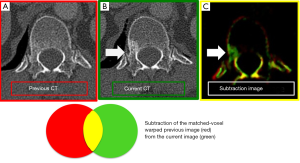
Non-rigid image registration methods have been proposed to reduce further subtraction artifacts on TS images (27-29). hereby we provide example of some of registration methods for reducing the subtraction artifacts due to normal structures, including blood vessels, in order to get precise subtraction of a pair of CT images. Figure 1 illustrates the overall scheme of a general 3D image registration method. In the first stage of the image registration method, normalization of the voxel size on the previous and current images is performed based on a linear interpolation method. In the second stage, global matching and local image matching techniques to adjust for the global and local displacement due to variation in positioning are performed. Finally, voxel matching technique or generalized gradient vector flow (GGVF) based matching technique for more accurate registration are performed as a 3D non-rigid image warping technique. Then, a TS image is obtained by subtraction from warped previous image to the current one. Figure 2 shows the notion of the registration scheme. An image warping technique such as voxel matching is applied to the current and the previous image (30), as shown in Figure 2A and Figure 2C, respectively, in order to get shift vectors, which represent the extent of deformation (or warping) of the previous image relative to the current image. These shift vectors on the current image will enable us to warp the previous image to produce a TS image. The voxel matching technique is an optimization non-rigid registration tools for reduction of the subtraction artifacts in a TS method. It can be affixed to 2D images including chest radiographs by replacing the voxel matching in 3D images with a pixel matching method.

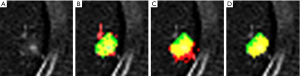
For the registration of bone on CT, some methods are used in order to reduce misregistration artifacts. Large Deformation Diffeomorphic Metric Mapping (LDDMM), which is included in non-rigid image registration technique, can handle copious deformation without losing the topology of the object (31-33). With LDDMM, the connected and disconnected structures keep that way, even with severe local transformation. The salient region feature (SRF) is more reliable method than the density based image registration method with respect to alignment and is unaffected by the local density change, and therefore, image matching based on SRF is suitable for bone subtraction images (34) (Figure 3).
Application for lesion detection on MDCT image based on TS technique
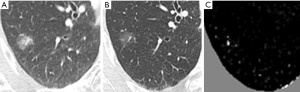
Misregistration artifacts still exist on the TS images after a non-rigid registration processing, and therefore, several methods are applied for the subtraction images to reduce them. Segmentation is another big issue in object detection in image processing. There are many approached such as Graph Cuts and SuperVoxel etc. 2D or 3D detection methods are widely used in medical imaging and video processing. The SuperVoxel is a segmentation method based on voxels with similar attributes. Graph Cuts is a graph-based segmentation technique. Both-are sometimes used to segment the subtle lesions such as ground-glass lung nodules in initial candidate region (35). When a temporal previous image is generated using the method of the previous study, large nodular opacity and its temporal change are highlighted on the TS image, however, changes in small ground-glass nodule disappear in some cases. An example is shown in Figure 4. A localized ground-glass nodule exists on both current and previous images, but the temporal change cannot be emphasized precisely because of lightness in the subtraction image. Because of this, it is considered to be the forcing power when the alignment process is high, which is intended to significantly reduce artifacts as a result of misalignment of the blood vessels and chest wall. A new method was developed to overcome this problem. Figure 5 demonstrates the outline of the proposed processing architecture. The whole processing is followed by the initial TS image generation, region-of-interest (ROI) extraction, Graph Cut-based ground-glass nodules 3D-ROI detection, support vector machine (SVM)-based shadow removal, followed by the coordinates of TS images to assist the voxel matching of previous image. Ultimately, the current image and the registered image was subtracted. Figure 6 shows a part of the extraction results by 2D display according to the 3 types of region extraction methods: (I) only Graph Cuts; (II) SuperVoxel and Graph Cuts; (III) SuperVoxel and Graph Cuts using the gradient vector concentration degree. Figure 7 displays TS images without voxel matching method (Global Matching + Local Matching + Elastic Matching) and with voxel matching (Global Matching + Local Matching + Elastic Matching + Voxel Matching), and the final subtraction image by the proposed method.

Clinical study based on TS technique
Several clinical studies using CT-TS technique indicate the improvement of diagnostic ability for lung nodule and bone metastasis detection (28,30,36-42). The performance for the detection of lung nodule and bone metastasis was summarized in Table 1. Although the technique to create CT-TS images is various, all the previous reports demonstrated the CT-TS improved the diagnostic performance of observers.
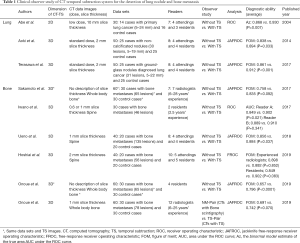
Full table
Lung nodule
Abe et al. first reported the clinical efficiency of CT-TS using 2D registration technique. This is the sole observer performance study for 2D-TS method. They performed ROC analysis using films of low dose CTs, which were obtained with a single-detector helical CT scanner using 10 mm collimation, and found that the TS technique can increase the sensitivity and specificity for detection of lung cancer (36). We developed 3D-CT-TS system using a voxel matching technique with GGVF algorithms and assessed the performance of TS on thoracic CT using a soft copy display (28). Value of conventional ROC analysis is limited because only one lesion can be used per case and the location of the lesion cannot be considered in the evaluation. By contrast, jackknife free-response receiver operating characteristics (JAFROC) analysis permits us to evaluate radiologists’ performance in diagnosing CT images by using multiple responses, each with information on the confidence level and location (3,43). As the results of JAFROC observer performance study including 30 non-calcified nodules less than 20 mm, the average figure-of-merit (FOM) values for all radiologists increased to a statistically significant degree, from 0.838 without the CT-TS images to 0.894 with the images while the reading time with CT-TS images was not significantly different from that without. In another observer performance study for the lung cancer with predominant ground-glass opacity (ranging from 5 to 22 mm), the use of CT-TS improved the diagnostic performance without considerably extending the reading time (37) (Figure 8). Despite the lack of an observer performance study, there are two other studies that evaluated the displacement of landmarks placed at the bifurcations of segmental bronchi/vessels (33,44). Takao et al. evaluated the registration accuracy based on the concept of target registration error using 19 landmarks chosen at the bifurcations of segmental bronchi, and indicated the average displacements of the landmarks were less than the dimensions of a voxel (44). Another study evaluated the registration accuracy of non-linear registration using LDDMM by measuring the displacement of landmarks placed on vessel bifurcations for each lung segment, and found the average displacement of landmarks after LDDMM transformation with cascading elasticity control was significantly smaller than that after affine transformation (33).
Bone metastasis
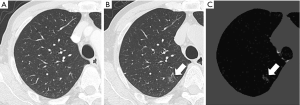
Several studies have reported the value of CT-TS for the detection of the bone metastases recently. Sakamoto et al. assessed the utility of the CT-TS method using LDDMM registration algorithm, and indicated that the average reading time to detect bone metastases on CT images shortened significantly with the CT-TS (38). The average FOM increased from 0.758 without CT-TS images to 0.835 with CT-TS images, although this difference was not statistically significant. Ueno et al. assessed the value of the CT-TS system with a final local matching technique based on SRF. As the results of the observer study including 20 pairs (current and previous CT images) with sclerotic bone metastasis in the thoracolumbar spine (135 lesions) and 20 pairs of those in patients without bone metastasis, this study indicated that the use of CT-TS improved the detectability of the sclerotic bone metastasis and shorten the reading time (30). There is only one prior comparative study dealing with observer performance between CT-TS method and the other modality. Onoue et al. compared the observer performance in detecting bone metastases of CT-TS using LDDMM with that of bone scintigraphy with planar scan and single-photon emission computed tomography (SPECT) (41). They used data on 60 patients (30 patients with bone metastasis and 30 patients without) and 12 readers participated in the observer performance study, The sensitivity of CT-TS was significantly higher than that of bone scintigraphy (54.3% vs. 41.3%) and the FOM of CT-TS appeared superior to that of bone scintigraphy (0.742 vs. 0.691). They concluded that CT-TS might be superior to bone scintigraphy including SPECT for early detection of bone metastasis.
Clinical implication
Following the previous observer performance results, CT-TS method seems to assist the radiologists’ interpretation for lung nodule and bone metastasis detection sufficiently. In general, less experienced observers benefited from CT-TS more, although the diagnostic ability for readers improved from with CT-TS regardless of experience. Nevertheless, the clinical usage of CT-TS system may be limited since both previous and current images are required. CT is routinely used among cancer patients to assess the therapeutic effect and to check for local recurrence and metastasis. Moreover, CT follow-up examinations in cancer patients are often repeated many times. Therefore, its use may be practical in follow-up examinations of the patients who have a high risk of lung and bone lesions such as patients with malignant tumor. On the other hand, small lung nodules and localized ground-glass lesions are being more readily detected with recent increased use of CT. CT-TS system may also be useful in repeated CT lung cancer screening.
Misregistration artifacts of CT-TS method, resulting from mismatching normal structures in current and previous images, lead misdiagnosis and extend the interpretation times. At this point, there are still open challenges to overcome the further reduction of the misregistration artifacts on CT-TS system. For clinical use, technical development that enables more precise registration is required.
Conclusions
We reviewed the CT-TS techniques and applications to detect lung and bone abnormalities. The CT-TS would improve the diagnostic performance of observer, and the applications to other lesions will be possible in the near future. When using CT-TS, the diagnostic performance is heavily affected by the image quality of the subtraction image. In order to generate CT-TS images more precisely, further development of the region extraction methods and applications is desired.
Acknowledgments
Funding: None.
Footnote
Provenance and Peer Review: With the arrangement by the Guest Editors and the editorial office, the article has undergone external peer review.
Conflicts of Interest: All authors have completed the ICMJE uniform disclosure form (available at http://dx.doi.org/10.21037/qims-20-1367). The authors have no conflicts of interest to declare.
Open Access Statement: This is an Open Access article distributed in accordance with the Creative Commons Attribution-NonCommercial-NoDerivs 4.0 International License (CC BY-NC-ND 4.0), which permits the non-commercial replication and distribution of the article with the strict proviso that no changes or edits are made and the original work is properly cited (including links to both the formal publication through the relevant DOI and the license). See: https://creativecommons.org/licenses/by-nc-nd/4.0/.
References
- Guo W, Li Q. High performance lung nodule detection schemes in CT using local and global information. Med Phys 2012;39:5157-68. [Crossref] [PubMed]
- Berlin L. Malpractice issues in radiology: perceptual errors. AJR Am J Roentgenol 1996;167:587-90. [Crossref] [PubMed]
- Sahiner B, Chan HP, Hadjiiski LM, Cascade PN, Kazerooni EA, Chughtai AR, Poopat C, Song T, Frank L, Stojanovska I, Attili A. Effect of CAD on radiologists’ detection of lung nodules on thoracic CT scans: Analysis of an observer performance study by nodule size. Acad Radiol 2009;16:1518-30. [Crossref] [PubMed]
- Kano A, Doi K, MacMahon H, Hassell DD, Giger ML. Digital image subtraction of temporally sequential chest images for detection of interval change. Med Phys 1994;21:453-61. [Crossref] [PubMed]
- Ishida T, Ashizawa K, Engelmann R, Katsuragawa S, MacMahon H, Doi K. Application of temporal subtraction for detection of interval changes on chest radiographs: Improvement of subtraction images using automated initial image matching J Digit Imaging 1999;12:77-86. [Crossref] [PubMed]
- Difazio MC, MacMahon H, Xu XW, Tsai P, Shiraishi J, Aramoto SG III, Doi K. Digital chest radiography: Effect of temporal subtraction images on detection accuracy. Radiology 1997;202:447-52. [Crossref] [PubMed]
- Uozumi T, Nakamura K, Watanabe H, Nakata H, Katsuragawa S, Doi K. ROC analysis of detection of metastatic pulmonary nodules on digital chest radiographs with temporal subtraction. Acad Radiol 2001;8:871-8. [Crossref] [PubMed]
- MacMahon H. Improvement in detection of pulmonary nodules: digital image processing and computer-aided diagnosis. Radiographics 2000;20:1169-77. [Crossref] [PubMed]
- Li Q, Li F, Doi K. Computerized detection of lung nodules in thin-section CT images by use of selective enhancement filters and an automated rule-based classifier. Acad Radiol 2008;15:165-75. [Crossref] [PubMed]
- Elsen PA van den, Pol EJD, Viergever MA. Medical image matching-a review with classification. Conf Proc IEEE Eng Med Biol 1993;12:26-39.
- Maintz JBA, Viergever MA. A survey of medical image registration. Med Image Anal 1998;2:1-36. [Crossref] [PubMed]
- Zitoba B, Flusser J. Image registration method: a survey. Image Vis Comput 2003;21:977-1000. [Crossref]
- Ko JP, Betke M. Chest CT: automated nodule detection and assessment of change over time--preliminary experience. Radiology 2001;218:267-73. [Crossref] [PubMed]
- Besl PJ, McKay ND. A method for registration of 3-D shapes IEEE Trans Pattern Anal Mach Intell 1992;14:239-56. [Crossref]
- Maintz JBA, Elsen PA, van den , Viergever MA. Comparison of edge-based and ridge-based registration of CT and MR brain images. Med Image Anal 1996;1:151-61. [Crossref] [PubMed]
- Lester H, Arridge SR. A survey of hierarchical non-linear medical image registration Pattern Recognit ;1999:129-49. [Crossref]
- Babaud J, Witkin AP, Baudin M, Duda RO. Uniqueness of the Gaussian kernel for scale-space filtering IEEE Trans Pattern Anal Mach Intell 1986;8:26-33. [Crossref] [PubMed]
- Haralick RM. SternBerg SR, Zhuang X. Image analysis using mathematical morphology. IEEE Trans Pattern Anal Mach Intell 1987;9:532-50. [Crossref] [PubMed]
- Unse M, Aldroubi A, Eden M. Fast B-spline transforms for continuous image representation and interpolation. IEEE Trans Pattern Anal Mach Intell 1991;13:277-85. [Crossref]
- Rueckert D, Sonoda LI, Hayes C, Hill DLG, Leach MO, Hawkes DJ. Nonrigid registration using free-from deformations: Application to breast MR images IEEE Trans Med Imaging 1999;18:712-21. [Crossref] [PubMed]
- Periaswamy S, Farid H. Elastic registration in the presence of intensity variations. IEEE Trans Med Imaging 2003;22:865-74. [Crossref] [PubMed]
- Pariaswamy S, Farid H. Medical image registration with partial Ddta. Med Imag Anal 2006;13:452-64. [Crossref]
- Walimbe V, Shekhar R. Automatic elastic image registration by interpolation of 3D rotations and translations from discrete rigid-body transformation Med Image Anal 2006;10:899-914. [Crossref] [PubMed]
- Wu G, Kim M, Wang Q, Munsell BC, Shen D. Scalable High-Performance Image Registration Framework by Unsupervised Deep Feature Representations Learning. IEEE Trans Biomed Eng 2016;63:1505-6. [Crossref] [PubMed]
- Hu Y, Modat M, Gibson E, Li W, Ghavami N, Bonmati E, Wang G, Bandula S, Moore CM, Emberton M, Ourselin S, Noble JA, Barratt DC, Vercauteren T. Weakly-supervised convolutional neural networks for multimodal image registration. Med Image Anal 2018;49:1-13. [Crossref] [PubMed]
- Eppenhof KAJ, Lafarge MW, Moeskops P, Veta M, Pluim JPW. Deformable image registration using convolutional neural networks. Medical Imaging 2018: Image Processing. International Society for Optics and Photonics, 2018;10574:105740S [PubMed]
- Itai Y, Kim H, Ishikawa S, Katsuragawa S, Doi K. Development of a voxel matching technique for substantial reduction of subtraction artifacts in temporal subtraction images obtained from thoracic MDCT. J Digit Imaging 2010;23:31-8. [Crossref] [PubMed]
- Aoki T, Murakami S, Kim H, Fujii M, Takahashi H, Oki H, Hayashida Y, Katsuragawa S, Shiraishi J, Korogi Y. Temporal subtraction method for lung nodule detection on successive thoracic CT soft-copy images. Radiology 2014;271:255-61. [Crossref] [PubMed]
- Yoshino Y, Miyajima T, Lu H, Tan JK, Kim H, Murakami S, Aoki T, Tachibana R, Hirano Y, Kido S. Automatic classification of lung nodules on MDCT images with the temporal subtraction technique. Int J Comput Assist Radiol Surg 2017;12:1789-98. [Crossref] [PubMed]
- Ueno M, Aoki T, Murakami S, Kim H, Terasawa T, Fujisaki A, Hayashida Y, Korogi Y. CT temporal subtraction method for detection of sclerotic bone metastasis in the thoracolumbar spine. Eur J Radiol 2018;107:54-9. [Crossref] [PubMed]
- Qiu A, Albert M, Younes L, Miller MI. Time sequence diffeomorphic metric mapping and parallel transport track time-dependent shape changes. Neuroimage 2009;45:S51-60. [Crossref] [PubMed]
- Glaunès J, Qiu A, Miller MI, Younes L. Large Deformation Diffeomorphic Metric Curve Mapping. Int J Comput Vis 2008;80:317-36. [Crossref] [PubMed]
- Sakamoto R, Mori S, Miller MI, Okada T, Togashi K. Detection of time-varying structures by large deformation diffeomorphic metric mapping to aid reading of high-resolution CT images of the lung. PLoS One 2014;9:e85580 [Crossref] [PubMed]
- Huang X, Sun Y, Metaxas D, Sauer F, Chenyang X. Hybrid image registration based on configual matching of scale invariant salient region features. Proc of the 2004 IEEE Computer Society Conference on Computer Vision and Pattern Recognition Workshop 2004:167-79.
- Boykov Y, Kolmogorov V. An experimental comparison of min-cut/max-flow algorithms for energy minimization in vision. IEEE Trans Pattern Anal Mach Intell 2004;26:1124-37. [Crossref] [PubMed]
- Abe H, Ishida T, Shiraishi J, Li F, Katsuragawa S, Sone S, Macmahon H, Doi K. Effect of temporal subtraction images on radiologists’ detection of lung cancer on CT: results of the observer performance study with use of film computed tomography images. Acad Radiol 2004;11:1337-43. [Crossref] [PubMed]
- Terasawa T, Aoki T, Murakami S, Kim H, Fujii M, Kobayashi M, Chihara C, Hayashida Y, Korogi Y. Detection of lung carcinoma with predominant ground-glass opacity on CT using temporal subtraction method. Eur Radiol 2018;28:1594-9. [Crossref] [PubMed]
- Sakamoto R, Yakami M, Fujimoto K, Nakagomi K, Kubo T, Emoto Y, Akasaka T, Aoyama G, Yamamoto H, Miller MI, Mori S, Togashi K. Temporal Subtraction of Serial CT Images with Large Deformation Diffeomorphic Metric Mapping in the Identification of Bone Metastases. Radiology 2017;285:629-39. [Crossref] [PubMed]
- Iwano S, Ito R, Umakoshi H, Karino T, Inoue T, Li Y, Naganawa S. Thoracic Temporal Subtraction Three Dimensional Computed Tomography (3D-CT): Screening for Vertebral Metastases of Primary Lung Cancers. PLoS One 2017;12:e0170309 [Crossref] [PubMed]
- Hoshiai S, Masumoto T, Hanaoka S, Nomura Y, Mori K, Hara T, Saida T, Okamoto Y, Minami M. Clinical usefulness of temporal subtraction CT in detecting vertebral bone metastases. Eur J Radiol 2019;118:175-80. [Crossref] [PubMed]
- Onoue K, Nishio M, Yakami M, Aoyama G, Nakagomi K, Iizuka Y, Kubo T, Emoto Y, Akasaka T, Satoh K, Yamamoto H, Isoda H, Togashi K. CT temporal subtraction improves early detection of bone metastases compared to SPECT. Eur Radiol 2019;29:5673-81. [Crossref] [PubMed]
- Onoue K, Nishio M, Yakami M, Sakamoto R, Aoyama G, Nakagomi K, Iizuka Y, Kubo T, Emoto Y, Akasaka T, Satoh K, Yamamoto H, Isoda H, Togashi K. Temporal subtraction of computed tomography images improves detectability of bone metastases by radiology residents. Eur Radiol 2019;29:6439-42. [Crossref] [PubMed]
- Hirose T, Nitta N, Shiraishi J, Nagatani Y, Takahashi M, Murata K. Evaluation of computer-aided diagnosis (CAD) software for the detection of lung nodules on multidetector row computed tomography (MDCT): JAFROC study for the improvement in radiologists' diagnostic accuracy. Acad Radiol 2008;15:1505-12. [Crossref] [PubMed]
- Takao H, Doi I, Tateno M. Evaluation of an automated system for temporal subtraction of thin-section thoracic CT. Br J Radiol 2007;80:85-9. [Crossref] [PubMed]




