
Identifying subtypes of mild cognitive impairment from healthy aging based on multiple cortical features combined with volumetric measurements of the hippocampal subfields
Introduction
Alzheimer’s disease (AD) is a progressive neurodegenerative disease characterized by neurofibrillary tangles (NFT), amyloid deposition (senile plaque, SP), and neuronal loss in the basal forebrain. AD is accompanied by comprehensive cognitive decline, behavioral changes, and dysfunction. The incidence rate of AD is 10–20% among individuals aged 65 or older (1); AD brings a heavy burden to patients, their families, and society (2). There is a pressing need to investigate the abnormal changes in the brains of patients in the early stages of symptomatic dementia with mild cognitive impairment (MCI) due to the irreversibility of AD, to offer opportunities for timely intervention. It is of great importance that the patients with MCI who are most likely to develop AD are identified to provide them with early pharmacological treatment and effective interventions.
Magnetic resonance imaging (MRI) technology provides an extremely intuitive and reliable basis for the diagnosis of neurodegenerative diseases. Several studies have confirmed that specific anatomical regions of MCI/AD patients are susceptible to specific neurodegenerative processes. In the early stage, patients suffer from functional neuronal disorders due to changes in cortical thinning and loss of gray matter (3-6). It is well known that the hippocampus region plays a vital role in the short- to long-term information integration process for memory. The hippocampus is most likely to be the first brain region to suffer damage.
Furthermore, clinical studies have shown that the hippocampus is one of the most effective and widely used biomarkers for revealing conversion from MCI to AD (7-9). However, it is usually viewed as a single entity because of the coarse resolution of MR images. With the substantial developments in high-resolution MRI data acquisition technology, new opportunities for explicitly exploring individual hippocampal subfields have emerged. It is now possible to explore the fimbria, presubiculum, subiculum, dentate gyrus (DG), hippocampus-amygdala-transition-area (HATA), the four cornu ammonis regions (CA1–CA4), and the hippocampus, hippocampal fissure, and hippocampal tail (10-12). It has been reported that CA1 measurements were more sensitive than global hippocampal volumetry for detecting structural changes at the pre-dementia stage (13,14). It was also found that hippocampal subfields were associated with age-related memory decline and distinct aspects of memory formation (12).
The cerebral cortical surfaces reflect the different neuropathological mechanisms of patients with MCI and AD. The cortical thickness, for example, reveals the degree of brain atrophy by characterizing the shortest distance between the inner and outer surfaces of the cerebral cortex. The surface area represents the degree of cortical folds, and the difference in mean curvature is caused by thickness variations or surface area changes that provide practical information on the folding patterns and volume changes in the cerebral cortex. These cortical surface properties may exhibit different developmental trajectories for different neurological diseases (15). These differences play a vital role in the diagnostic classification of MCI and AD, and in the early identification of MCI patients who carry the risk of later converting to AD (16,17).
In this study, MR images of patients with MCI and NC were collected. The FreeSurfer package (18) was applied to segment brain tissue automatically and calculate cortical features (surface area, gray matter volume, cortical thickness, and mean curvature) of 68 brain regions. Following that, we extracted the voxel characteristics of 16 hippocampal subfields. Then an analysis of variance (ANOVA) model was used to reveal the significant differences between the compared groups. Two types of features, including cortical surface measurements and volume of hippocampal subfields, were combined and used as the input of the support vector machine-recursive feature elimination (SVM-RFE) for discriminative analysis. The most discriminative features corresponding to the brain regions were extracted to elucidate the path of brain structural deterioration and cognitive dysfunction in patients with MCI to achieve early detection and conversion risk assessment of MCI.
Methods
Subjects
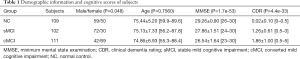
Included in this study were 322 subjects selected from the Alzheimer’s Disease Neuroimaging Initiative (ADNI) database (http://adni.loni.usc.edu/). This study included 109 NC, 102 stable MCI (sMCI), and 111 converted MCI (cMCI) subjects. The MR imaging data for each subject were acquired and each subject was evaluated using two cognitive impairment measurements: the mini-mental state examination (MMSE) and the clinical dementia rating (CDR). Next, the data were analyzed at three distinct time points: baseline, 12 months, and 24 months. Among the identified MCI patients at baseline, an individual was considered as cMCI if they had converted to AD within two years; the rest were considered as sMCI subjects. Table 1 shows the subject demographic information and dementia status.

Full table
Image acquisition
The datasets included standard T1-weighted MR images. The images were obtained using volumetric 3D MPRAGE (magnetization prepared rapid gradient echo) 1.5 T scanners with a 1.25×1.25-mm in-plane spatial resolution and 1.2-mm-thick sagittal slices, with a pixel resolution of 256×256. The data were collected from a variety of scanners using the protocols specified by the ADNI website (TR =2 s, TE =2.63 ms, FOV =25.6 cm). All data were preprocessed with intensity normalizing and gradient unwarping.
Age, MMSE, and CDR had significant differences among the three comparison groups using the ANOVA model. Sex also had significant differences among the three groups using the Chi-square test.
Image processing and features calculation
The images were processed and analyzed by FreeSurfer software version 5.0.0 (http://surfer.nmr.mgh.harvard.edu/), which provides a set of tools that can be used to analyze and visualize structural and functional brain images. The FreeSurfer pipeline includes a series of processing steps such as head motion correction, inhomogeneous field correction, removal of non-brain tissue, among many others. The pipeline is then able to produce regional cortical thickness and volumetric measurements, including the local curvature, surface area, and the surface normal. According to the Desikan-Killiany atlas (19), the left and right hemispheres were divided into 34 brain regions simultaneously, and thus 68 cortical measurements corresponding to the brain regions were obtained. Four measurements, including the cortical thickness, gray matter volume, surface area, and average curvature, were considered in this study. Furthermore, the individual hippocampal subfields were acquired based on the parcellation scheme (11).
Hippocampal subfields delineation
The FreeSurfer algorithm employed Bayesian inferences from prior probabilistic atlases built with ultra-high resolution ex vivo MRI data (~0.1 mm isotropic) to obtain an automated segmentation of the hippocampal substructures that had shown strong correlations with manual delineations (11,20,21). Following the segmentation scheme presented in (10), eight subfields were extracted in each hemisphere including the fimbria, CA1, CA2_3, CA4_DG, presubiculum, subiculum, hippocampal fissure, the hippocampus. CA2 and CA3 were combined because of unclear contrast. Next, the alveus volume was removed because of the thin shape and unreliable segmentation (11,12); thus, 16 subfields were extracted by the FreeSurfer 5.3.0.
Statistics analysis
ANOVA is a statistical method used to evaluate differences between two or more means. The F-distribution function is adopted to analyze the variances from each population and groupings of populations. The analysis determined whether the variability between and within each population was significantly different, and also determined the influence that the independent variables had on the dependent variable in a regression study. Bonferroni correction was applied to correct the increased error rates in hypothesis testing that had multiple comparisons.
The Chi-square and ANOVA tests were performed on sex and age to determine whether there were significant differences between the groups. The ANOVA model with subsequent Bonferroni correction was applied to investigate the differences in cortical measurements and volumes of hippocampal subfields among the three groups.
Classifier
SVMs are a set of supervised learning methods based on the statistical learning theory and structural risk minimization. These methods have been widely used in non-linear classification, regression, function estimation, and density estimation, among many others. Since the dimension of the features was less than the sample size, a linear kernel was selected in the SVM classifier following a suggestion given by the LIBSVM toolbox (https://www.csie.ntu.edu.tw/~cjlin/libsvm). After that, owing to the small sample size, a 10-fold cross-validation method was selected. The RFE strategy (16) was adopted to achieve the best performance and explore the dominant features associated with core regions. The RFE explored the algorithm to remove unimportant or redundant features one-by-one based on the assigned classification weight during classification to perform feature dimension reduction. Since the weight of each feature is represented by its discriminative ability, the retained features could be used with the SVM classifier to achieve promising performance.
The SVM-RFE classification of the three groups was performed using the cortical features and hippocampal subfields, and the performance was evaluated using the accuracy (Acc), sensitivity (Sen), specificity (Spe), and area under the curve (AUC). The receiver operating characteristic (ROC) curve was defined to assess the performance of a classifier in decision-making. The ROC curve illustrates the trade-off of true-positives versus the false-positives as the discriminating threshold varied from 0 to 1. In a ROC curve, the true-positive rate (sensitivity) was plotted in the function of the false-positive rate (100-specificity) for the different cut-off points of each threshold. Each point on the ROC curve represents a sensitivity/specificity pair corresponding to a particular decision threshold. It is a measurement of how well a parameter can distinguish between the two diagnostic groups.
The area under a ROC curve (AUC) was then calculated by adding successive trapezoid areas below the ROC curve as an overall measurement of the performance of the classifier.
The criteria for determining the best feature dimension was that the SVM-RFE should achieve the best overall performance with a balance between accuracy, sensitivity, and specificity. The primary measurement considered was the accuracy, followed by the sensitivity, and then the specificity.
Results
Differences in cortical features
Table 2 illustrates that there were significant differences in most of the regions with bilateral symmetry in gray volume and cortical thickness between the three compared groups. Also, each region increased with significant differences from NC to sMCI, and then to NC-MCI, which meant that atrophies existed in many brain regions of the patients with sMCI. Patients with cMCI experienced more severe atrophies than patients with sMCI. However, only a few brain regions had significant differences in their surface area and average curvature between the compared groups.
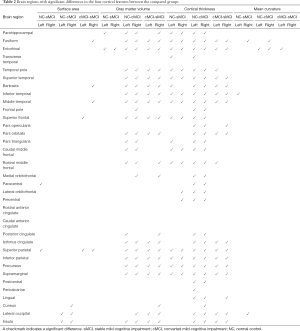
Full table
The cortical thickness showed significant differences in 33 brain regions, mainly located in the bilateral medial temporal lobe, left lateral temporal lobe, partial frontal lobe and partial parietal lobe for the NC-sMCI comparison as shown in Figure 1. Also, there existed significant differences in gray matter volume of the left parahippocampal gyrus, bilateral entorhinal, in mean curvature of three brain regions including the left inferior temporal, the right fusiform, and the right lateral occipital, and in surface area of two brain regions including the left paracentral, and the left superior parietal. For the cMCI-NC comparison, the cortical thickness in almost all of the brain regions had significant differences, as well as gray matter volume. However, statistical differences existed in only a couple of regions concerning the surface area and mean curvature.
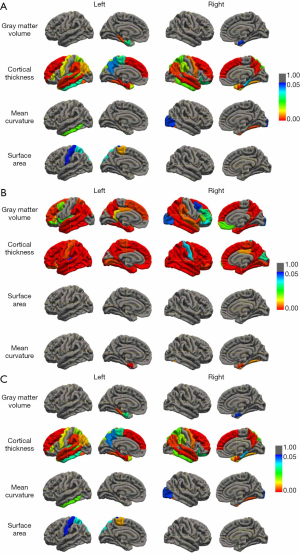
In the sMCI-cMCI comparison group, gray matter volume and cortical thickness had significant differences in most of the brain regions, mainly located in the temporal lobe, parietal lobe, occipital lobe, and frontal lobe, while only a few brain regions had statistical differences in the mean curvature and surface area.
Differences in hippocampal subfields
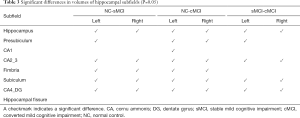
Table 3 illustrates that the volumes were significantly different in most subfields between the sMCI and NC groups, except for the bilateral CA1, hippocampal fissure, and right presubiculum. Similarly, the cMCI group had significant differences in the volumes in most of the subfields, except for the right CA1, and bilateral hippocampal fissure. Compared with the sMCI group, nine subfields had significant differences in volumes in the cMCI group, including the bilateral hippocampus, CA2_3, subiculum, CA4_DG, and left presubiculum.

Full table
Classification results
Each subject had 68 cortical features and 16 hippocampal subfield volumes. For the sMCI-NC comparison using the cortical thickness, 58 features were determined to achieve the best performance. The 62, 45, and 43 dominant features were retained using gray matter volume, mean curvature, and surface area, respectively. The SVM-RFE classifier obtained its best performance when the 20, 44, 48, and 61 cortical features were selected for the cMCI-NC discrimination using cortical thickness, gray matter volume, mean curvature, and surface area, respectively. At the same time, the 63, 63, 43, and 35 features had a prominent ability to classify using the four types of cortical features. The volumes of the 2, 5, and 13 hippocampal subfields were extracted for the sMCI-NC, cMCI-NC, and sMCI-cMCI comparisons, respectively. Table 4 lists the classification performance in detail.
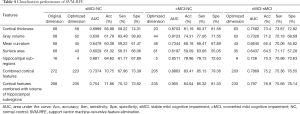
Full table
Discussion
This study focuses on the discriminative analysis of cortical surface measurements and the volume of hippocampal subfields among NC, sMCI, and cMCI groups. The SVM-REF classifier was adopted to perform classification as a model that integrated feature reduction with classification.
Experimental results suggested that the SVM-RFE classifier acquired better performance for sMCI-NC discrimination when four types of cortical features were considered. A combination of the four types of cortical features and the volumes of the hippocampal subfields exhibited the best power, with an accuracy of 70.8%, sensitivity of 68%, specificity of 73.4%, and AUC of 0.75, respectively. The classifier obtained the highest accuracy (84.6%), sensitivity (85.3%), specificity (81.3%), and AUC (0.91) in the cMCI-NC comparison. The best performance in the cMCI-sMCI discrimination was reached with an accuracy of 76.9%, sensitivity of 75.7%, specificity of 78.1%, and AUC of 0.8. These results illustrate that the joint features could significantly improve the discriminative performance of the SVM-REF classifier.
A recent study (22) reported similar performances with an overall accuracy of 75% and AUC of 0.76 in discrimination of NC, early MCI, late MCI, and AD. Cortical surface morphometry of medial temporal lobe structures, such as the hippocampus and entorhinal cortex, were superior to the volumetric assessment of predicting the conversion from MCI to AD (23). Recent MRI studies have also investigated classification in MCI, AD patients, and aging normal individuals (24), and estimated their MCI conversion risk (25).
The longitudinal exploration of classifying NC, MCI, and AD and conversion prediction has also attracted attention in numerous other recent studies. It has been shown that the parahippocampal gyrus and fusiform gyrus volumes were significantly reduced in the cMCI group compared to the sMCI group and obtained an accuracy of 0.779 in conversion prediction (26). The combination of the hippocampus and entorhinal cortex yielded a significantly higher AUC in the 5-year follow-up (0.73 at 2 years vs. 0.84 at 5 years) (27). Based on hippocampal volume, the volume of 47 cortical and subcortical regions, ensemble linear discriminant models acquired prediction accuracy of 63.8% to 77.0% of future conversion from MCI to AD at 6, 12, 24, 36, and 48 months.
It was observed that the subtypes of MCI had atrophy or experienced cortical thinning widely in the regions of the lateral temporal cortex, frontal cortex, parietal cortex, and olfactory cortex concerning the relationship between the required regional distribution and neuropsychological dysfunction. These results were especially true regarding the gray matter volume, and the cortical thickness exhibited a significantly different structural pattern in many regions (Table 2). With the conversion from sMCI to AD, it can be seen that the sMCI group experienced further decline in the regions of the temporal gyrus, olfactory cortex, and left temporal pole. Meanwhile, the cMCI patients had severe abnormal changes in most of the regions (Table 2 and Figure 1). As reported in our previous longitudinal study (28), these regions were highly correlated with emotion processing, learning, memory, decision-making, sense, recognition, and vision.
Many studies have reported similar findings in AD exploration, Venkatraghavan et al. (29) applied the event-based models (EBM) to exploit high-dimensional voxel-wise imaging biomarkers based on semi-supervised SVM to estimate temporal ordering of neuropathological changes in the brain structure using cross-sectional data from 1,737 subjects from the ADNI dataset. It was found that the atrophies with the progression to AD mainly occurred in 15 brain regions, including the bilateral medial and inferior temporal gyri, posterior temporal lobe, anterior temporal lobe, parahippocampal et ambiances, lateral occipitotemporal gyrus, hippocampus, amygdala, and right insula. In another study (30) including MRI scans from 699 subjects (NC, MCI, and AD), 259 cortical features of 8 types of regional measurements were extracted to perform multivariate analysis as well as SVM classification. Variables with the greatest importance for the separation between NC and AD were located in medial temporal lobe structures such as the hippocampus, amygdala, and entorhinal cortex. The combination of cortical thickness measurements with subcortical volumes showed potential for separating AD subjects from cognitively normal subjects to prospectively predict future conversion to AD from baseline. However, hippocampal subfields have not yet been fully explored.
The hippocampus belongs to the limbic system and plays important roles in the consolidation of information from short- to long-term memory, and in spatial memory, it enables navigation. Human studies demonstrated that entorhinal and CA1 were among the first regions to show preclinical AD pathology (31,32); focal atrophy in CA1 was also found by in vivo imaging studies of patients with both MCI and mild AD (33). CA1 was validated to be associated with verbal and visual episodic memory (34). The subiculum lies between the entorhinal cortex and the CA1 subfield of the hippocampus proper, which mainly receives input from CA1 and entorhinal cortical layer III pyramidal neurons and is the main output of the hippocampus. The pyramidal neurons in the subiculum exhibit transitions between two modes of action potential output: bursting and single spiking (35). The transitions between these two modes are thought to be important for routing information out of the hippocampus. The subiculum forms part of the cortical input to the entorhinal-hippocampal spatial and memory system, and it is associated with verbal and visual episodic memory. The fimbria, as a fiber bundle, covers the temporal parts of the hippocampus, and its stimulation leads to cholinergic excitation of CA1 oriens lacunosum-moleculare (OLM) cells (36). Compared with the NC group, the patients with sMCI, as well as the patients with cMCI, suffered abnormal changes in most of the subfields, leading to functional impairments in memory, spatial navigation, and control of attention. For the cMCI and sMCI comparison, there were significant differences in the bilateral presubiculum, subiculum, CA2_3, and CA4, covered from the surface layer to the deep layer. Therefore, the cMCI group experienced further damage to both signal pathways and functional fields, leading to serious degradation in memory, spatial perception, and attention. Izzo et al. (37) suggested that subicular hippocampal fields were most predictive of AD diagnosis. In particular, subicular and hippocampal fissure volume measurements might be effective in determining participants with MCI who are most likely to progress to AD. Several neuropathological studies have shown that the hippocampal subfields are differentially vulnerable to AD (13,38,39), and the annualized rate of CA1-3 atrophy was associated with an increased risk of developing Alzheimer’s clinical syndrome over time (40). Beyond these findings, our results illustrated that there were abnormal changes in most of the hippocampal subfields in the sMCI and cMCI groups, and the volumes of 13 subfields in particular showed significant differences between the sMCI and cMCI groups.
Our results show that cortical surface measurements can be used to assess global changes in brain structure, while hippocampal subfield volumetry can unveil local atrophy of the hippocampus. Therefore, their combination exhibits competitive performance in discriminating among the NC, sMCI, and cMCI groups. These findings could be greatly beneficial for explaining abnormal changes in brain structure and functional impairment, as well as early detection of AD and risk estimation of MCI conversion.
There are several limitations to this study. First, while we only used structural MR images, biomarkers such as FDG PET, PET-tau images, functional MR images, β-amyloid (Aβ), and apolipoprotein E (APOE) genotype could provide complementary information for detecting early AD and estimating the conversion risk of MCI. Second, only 322 subjects (102 sMCI, 111 cMCI, and 109 NC) were included in this study. With an increased sample size, a training set and independent testing set could have been established to validate the ability of the classifier and improve its generalization. Third, longitudinal analysis may have allowed the changes in biomarkers to be traced and effective and sensitive measurements in classification to be determined, especially in MCI progression. Fourth, more machine learning models, including random forest, XGBoost, and deep learning, should be considered and compared comprehensively in further study.
Conclusions
Based on MR brain images, this study investigated the differences in cortical features, including surface area, gray matter volume, cortical thickness, mean curvature, as well as hippocampal subfield volumes, between healthy older adults and patients with stable and cMCI. The SVM-RFE classifier was adopted to reduce dimensions and determine essential features for discrimination. With the progression from NC to sMCI and then to cMCI, the experimental results illustrated that the brain structure of subjects experienced progressive atrophic changes, which occurred in cortical measurements, especially in cortex thickness, gray matter volume, and hippocampus subfields. Compared with the single cortical features, the combination of four types of cortical measurements could achieve better results, and the volumes of hippocampal subfields could improve classification performance further. The regions with distinctive abilities were mainly located in the temporal, frontal, parietal, olfactory cortexes, and most of the hippocampal subfields, which offers essential evidence for clinical symptoms of cognitive and memory impairment in MCI patients. The cortical measurements, as well as volumes of hippocampal subfields, provided useful information for predicting and assessing the risk of MCI and its conversion.
Acknowledgments
Funding: This study was partly supported by Guangzhou Municipal Science and Technology (201604020170).
Footnote
Conflicts of Interest: All authors have completed the ICMJE uniform disclosure form (available at http://dx.doi.org/10.21037/qims-19-872). The authors have no conflicts of interest to declare.
Ethical Statement: The data used in this study were obtained from the Alzheimer’s Disease Neuroimaging Initiative (ADNI) (http://adni.loni.usc.edu/). The ADNI data were previously collected from across 50 research sites. Study subjects gave written informed consent at the time of enrolment for imaging and genetic sample collection and completed questionnaires approved by each participating sites’ Institutional Review Board (IRB). All procedures in studies involving human participants were performed following the ethical standards of the institutional and/or national research committee and the 1964 Helsinki declaration and its later amendments, or comparable ethical standards.
Open Access Statement: This is an Open Access article distributed in accordance with the Creative Commons Attribution-NonCommercial-NoDerivs 4.0 International License (CC BY-NC-ND 4.0), which permits the non-commercial replication and distribution of the article with the strict proviso that no changes or edits are made and the original work is properly cited (including links to both the formal publication through the relevant DOI and the license). See: https://creativecommons.org/licenses/by-nc-nd/4.0/.
References
- Talmelli LF, Gratao AC, Kusumota L, Rodrigues RA. Functional independence level and cognitive deficit in elderly individuals with Alzheimer's disease. Rev Esc Enferm USP 2010;44:933-9. [Crossref] [PubMed]
- Vos SJ, Xiong C, Visser PJ, Jasielec MS, Hassenstab J, Grant EA, Cairns NJ, Morris JC, Holtzman DM, Fagan AM. Preclinical Alzheimer's disease and its outcome: a longitudinal cohort study. Lancet Neurol 2013;12:957-65. [Crossref] [PubMed]
- Park M, Moon WJ, Structural MR. Imaging in the Diagnosis of Alzheimer's Disease and Other Neurodegenerative Dementia: Current Imaging Approach and Future Perspectives. Korean J Radiol 2016;17:827-45. [Crossref] [PubMed]
- Rummel C, Aschwanden F, McKinley R, Wagner F, Salmen A, Chan A, Wiest R. A Fully Automated Pipeline for Normative Atrophy in Patients with Neurodegenerative Disease. Front Neurol 2018;8:727. [Crossref] [PubMed]
- Ma X, Li Z, Jing B, Liu H, Li D, Li H. Identify the Atrophy of Alzheimer's Disease, Mild Cognitive Impairment and Normal Aging Using Morphometric MRI Analysis. Front Aging Neurosci 2016;8:243. [Crossref] [PubMed]
- Wei H, Kong M, Zhang C, Guan L, Ba M. for Alzheimer’s Disease Neuroimaging Initiative. The structural MRI markers and cognitive decline in prodromal Alzheimer's disease: a 2-year longitudinal study. Quant Imaging Med Surg 2018;8:1004-19. [Crossref] [PubMed]
- Platero C, Lin L, Tobar MC. Longitudinal Neuroimaging Hippocampal Markers for Diagnosing Alzheimer's Disease. Neuroinformatics 2019;17:43-61. [Crossref] [PubMed]
- Schröder J, Pantel J. Neuroimaging of hippocampal atrophy in early recognition of Alzheimer's disease--a critical appraisal after two decades of research. Psychiatry Res Neuroimaging 2016;247:71-8. [Crossref] [PubMed]
- Gaugler JE, Kane RL, Johnston JA, Sarsour K. Sensitivity and specificity of diagnostic accuracy in Alzheimer's disease: a synthesis of existing evidence. Am J Alzheimers Dis Other Demen 2013;28:337-47. [Crossref] [PubMed]
- Van Leemput K, Bakkour A, Benner T, Wiggins G, Wald LL, Augustinack J, Dickerson BC, Golland P, Fischl B. Automated Segmentation of Hippocampal Subfields From Ultra-High Resolution In Vivo MRI. Hippocampus 2009;19:549-57. [Crossref] [PubMed]
- Iglesias JE, Augustinack JC, Nguyen K, Player CM, Player A, Wright M, Roy N, Frosch MP, McKee AC, Wald LL, Fischl B, Van Leemput K. Alzheimer's Disease Neuroimaging Initiative. A computational atlas of the hippocampal formation using ex vivo, ultra-high resolution MRI: Application to adaptive segmentation of in vivo MRI. Neuroimage 2015;115:117-37. [Crossref] [PubMed]
- Zheng F, Cui D, Zhang L, Zhang S, Zhao Y, Liu X, Liu C, Li Z, Zhang D, Shi L, Liu Z, Hou K, Lu W, Yin T, Qiu J. The Volume of Hippocampal Subfields in Relation to Decline of Memory Recall Across the Adult Lifespan. Front Aging Neurosci 2018;10:320. [Crossref] [PubMed]
- La Joie R, Perrotin A, de La Sayette V, Egret S, Doeuvre L, Belliard S, Eustache F, Desgranges B, Chetelat G. Hippocampal subfield volumetry in mild cognitive impairment, Alzheimer's disease and semantic dementia. Neuroimage Clin 2013;3:155-62. [Crossref] [PubMed]
- La Joie R, Fouquet M, Mezenge F, Landeau B, Villain N, Mevel K, Pelerin A, Eustache F, Desgranges B, Chetelat G. Differential effect of age on hippocampal subfields assessed using a new high-resolution 3T MR sequence. Neuroimage 2010;53:506-14. [Crossref] [PubMed]
- Storsve AB, Fjell AM, Tamnes CK, Westlye LT, Overbye K, Aasland HW, Walhovd KB. Differential longitudinal changes in cortical thickness, surface area and volume across the adult life span: regions of accelerating and decelerating change. J Neurosci 2014;34:8488-98. [Crossref] [PubMed]
- Nesteruk M, Nesteruk T, Styczynska M, Barczak A, Mandecka M, Walecki J, Barcikowska-Kotowicz M. Predicting the conversion of mild cognitive impairment to Alzheimer's disease based on the volumetric measurements of the selected brain structures in magnetic resonance imaging. Neurologia I Neurochirurgia Polska 2015;49:349-53. [Crossref] [PubMed]
- Wu C, Guo SW, Hong YJ, Xiao BH, Wu YP, Zhang Q, Initia ADN. Discrimination and conversion prediction of mild cognitive impairment using convolutional neural networks. Quant Imaging Med Surg 2018;8:992-1003. [Crossref] [PubMed]
- Fischl B. FreeSurfer. Neuroimage 2012;62:774-81. [Crossref] [PubMed]
- Desikan RS, Ségonne F, Fischl B, Quinn BT, Dickerson BC, Blacker D, Buckner RL, Dale AM, Maguire RP, Hyman BT, Albert MS, Killiany RJ. An automated labeling system for subdividing the human cerebral cortex on MRI scans into gyral based regions of interest. Neuroimage 2006;31:968-80. [Crossref] [PubMed]
- Saygin ZM, Kliemann D, Iglesias JE, van der Kouwe AJW, Boyd E, Reuter M, Stevens A, Van Leemput K, McKee A, Frosch MP, Fischl B, Augustinack JC. Alzheimer's Disease Neuroimaging Initiative. High-resolution magnetic resonance imaging reveals nuclei of the human amygdala: manual segmentation to automatic atlas. Neuroimage 2017;155:370-82. [Crossref] [PubMed]
- Iglesias JE, Van Leemput K, Augustinack J, Insausti R, Fischl B, Reuter M. Alzheimer's Disease Neuroimaging Initiative. Bayesian longitudinal segmentation of hippocampal substructures in brain MRI using subject-specific atlases. Neuroimage 2016;141:542-55. [Crossref] [PubMed]
- Subramanyam Rallabandi VP, Tulpule K, Gattu M. Automatic classification of cognitively normal, mild cognitive impairment and Alzheimer's disease using structural MRI analysis. Informatics in Medicine Unlocked 2020;18:100305. [Crossref]
- Devanand DP, Bansal R, Liu J, Hao XJ, Pradhaban G, Peterson BS. MRI hippocampal and entorhinal cortex mapping in predicting conversion to Alzheimer's disease. Neuroimage 2012;60:1622-9. [Crossref] [PubMed]
- Basaia S, Agosta F, Wagner L, Canu E, Magnani G, Santangelo R, Filippi M. Alzheimer's Disease Neuroimaging Initiative. Automated classification of Alzheimer's disease and mild cognitive impairment using a single MRI and deep neural networks. Neuroimage Clin 2019;21:101645. [Crossref] [PubMed]
- Luk CC, Ishaque A, Khan M, Ta D, Chenji S, Yang YH, Eurich D, Kalra S. Alzheimer's Disease Neuroimaging Initiative. Alzheimer's disease: 3-Dimensional MRI texture for prediction of conversion from mild cognitive impairment. Alzheimers Dement (Amst) 2018;10:755-63. [Crossref] [PubMed]
- Mitolo M, Stanzani-Maserati M, Capellari S, Testa C, Rucci P, Poda R, Oppi F, Gallassi R, Sambati L, Rizzo G, Parchi P, Evangelista S, Talozzi L, Tonon C, Lodi R, Liguori R. Predicting Conversion From Mild Cognitive Impairment to Alzheimer's Disease Using Brain 1 H-MRS and Volumetric Changes: A Two- Year Retrospective Follow-Up Study. Neuroimage Clin 2019;23:101843. [Crossref] [PubMed]
- Moscoso A, Silva-Rodriguez J, Aldrey JM, Cortes J, Fernandez-Ferreiro A, Gomez-Lado N, Ruibal A, Aguiara P. Alzheimer's Disease Neuroimaging Initiative. Prediction of Alzheimer's disease dementia with MRI beyond the short-term: Implications for the design of predictive models. Neuroimage Clin 2019;23:101837. [Crossref] [PubMed]
- Guo S, Lai C, Wu C, Cen G. Conversion Discriminative Analysis on Mild Cognitive Impairment Using Multiple Cortical Features from MR Images. Front Aging Neurosci 2017;9:146. [Crossref] [PubMed]
- Venkatraghavan V, Dubost F, Bron EE, Niessen WJ, de Bruijne M, Klein S, Neuroimaging AD. Event-Based Modeling with High-Dimensional Imaging Biomarkers for Estimating Spatial Progression of Dementia. Information Processing in Medical Imaging 2019;11492:169-80. [Crossref]
- Westman E, Aguilar C, Muehlboeck JS, Simmons A. Regional Magnetic Resonance Imaging Measures for Multivariate Analysis in Alzheimer's Disease and Mild Cognitive Impairment. Brain Topogr 2013;26:9-23. [Crossref] [PubMed]
- Braak E, Braak H. Alzheimer's disease: transiently developing dendritic changes in pyramidal cells of sector CA1 of the Ammon's horn. Acta Neuropathol 1997;93:323-5. [Crossref] [PubMed]
- Braak H, Alafuzoff I, Arzberger T, Kretzschmar H, Del Tredici K. Staging of Alzheimer disease-associated neurofibrillary pathology using paraffin sections and immunocytochemistry. Acta Neuropathol 2006;112:389-404. [Crossref] [PubMed]
- Mueller SG, Schuff N, Yaffe K, Madison C, Miller B, Weiner MW. Hippocampal atrophy patterns in mild cognitive impairment and Alzheimer's disease. Hum Brain Mapp 2010;31:1339-47. [Crossref] [PubMed]
- Zammit AR, Ezzati A, Zimmerman ME, Lipton RB, Lipton ML, Katz MJ. Roles of hippocampal subfields in verbal and visual episodic memory. Behav Brain Res 2017;317:157-62. [Crossref] [PubMed]
- Cooper DC, Chung S, Spruston N. Output-mode transitions are controlled by prolonged inactivation of sodium channels in pyramidal neurons of subiculum. PLoS Biol 2005;3:e175. [Crossref] [PubMed]
- Leão RN, Mikulovic S, Leao KE, Munguba H, Gezelius H, Enjin A, Patra K, Eriksson A, Loew LM, Tort AB, Kullander K. OLM interneurons differentially modulate CA3 and entorhinal inputs to hippocampal CA1 neurons. Nat Neurosci 2012;15:1524-30. [Crossref] [PubMed]
- Izzo J, Andreassen OA, Westlye LT, van der Meer D. The association between hippocampal subfield volumes in mild cognitive impairment and conversion to Alzheimer's disease. Brain Res 2020;1728:146591. [Crossref] [PubMed]
- Wang L, Miller JP, Gado MH, McKeel DW, Rothermich M, Miller MI, Morris JC, Csernansky JG. Abnormalities of hippocampal surface structure in very mild dementia of the Alzheimer type. Neuroimage 2006;30:52-60. [Crossref] [PubMed]
- de Flores R, La Joie R, Chetelat G. Structural Imaging of Hippocampal Subfields in Healthy Aging and Alzheimer's Disease. Neuroscience 2015;309:29-50. [Crossref] [PubMed]
- Nadal L, Coupe P, Helmer C, Manjon JV, Amieva H, Tison F, Dartigues JF, Catheline G, Planche V. Differential annualized rates of hippocampal subfields atrophy in aging and future Alzheimer's clinical syndrome. Neurobiol Aging 2020;90:75-83. [Crossref] [PubMed]


